- Blog
- Live in london birdy shelter
- Trade show booth design plan
- Scipy stats linregress
- Dosbox windows 3-1 mouse feels weird
- Bollywood movie watch online
- Movavi video editor 11 key
- Tony hawk proving ground wii gameplay
- Smaart live calculate reverb time
- Show me how to stack g code soft shell mag pouches
- Zz top greatest hits lyrics
- Best beef protein powder brands
- Vuescan 9 serial
- Pokemon emerald emulator codes not working
- Bulgarian makarov pistol 8 round magazine
- Sget extension to surpac
#Scipy stats linregress license#
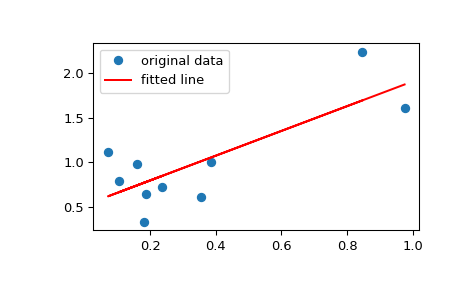
See the License for information about copying. This does not mean this is the best model for the data, just that the model fits well.Ĭopyright (C) 2013 by John Kitchin. zero is not included in any of the parameter confidence intervals. All of the parameters appear to be significant, i.e. Plt.savefig( 'images/linregress-conf.png')Ī fourth order polynomial fits the data well, with a good R^2 value. Plt.plot(time, Ca, 'bo', label= 'raw data') # plot fit import matplotlib.pyplot as plt Print ']'.format(beta - ci, beta + ci, beta) ST = t.ppf(1.0 - alpha/2.0, n - k) # student T multiplier Se = np.sqrt(np.diag(C)) # standard errorĪlpha = 0.05 # 100*(1 - alpha) confidence level Sigma2 = np.sum((Ca - np.dot(T, p))**2) / (n - k) # RMSEĬ = sigma2 * np.linalg.inv(np.dot(T.T, T)) # covariance matrix # the parameters are now in p # compute the confidence intervals Two-sided p-value for a hypothesis test whose null hypothesis is that the slope is zero, using Wald Test with t-distribution of the test statistic.Time = np.array()Ĭa = np.array()*1e-3 Would you be able to recommend any other mathematical statistics textbook?ĭo you think it's a good idea to rephrase the documentation to include this information about test statistic used? This is exactly what I've been seeing in the graphs attached in the post above we reject the null hypothesis more often than a given p-value/type-1 error threshold, like 0.05. Using the normal distribution in small samples should usually result in over-rejection, i.e. Regarding your "increased type 1 error" comment: This has already been a question since yesterday, but it hasn't attracted much attention.
I can vaguely recall seeing an argument for the t-distribution being the exact distribution if the residuals are normally distributed. estimates the regression for a polynomial of a single variable, but doesn't return much in terms of extra statisics.
#Scipy stats linregress code#
Using t-distribution for linear regression is "standard textbook" result, but I never read anything by Wasserman. only handles the case of a single explanatory variable with specialized code and calculates a few extra statistics. In many cases we have only the asymptotic theory with the normal distribution, but in practice in many cases wald test based on t distribution has better size in small samples than based on the normal distribution. Related aside: statsmodels allow users to choose between t and normal distribution (or F and chisquare for joint hypotheses). we reject the null hypothesis more often than a given p-value/type-1 error threshold, like 0.05.įor more information this would be a stats.stackexchange question. Normal and t distribution are the same asymptotically as the sample size increases, so both give the same answer in large samples, but using the normal distribution in small samples should usually result in over-rejection, i.e. Even when they are not normally distributed, then the t-distribution with the degrees of freedom correction is a better approximation to the true distribution of the test statistic in small samples. The t-distribution is the exact (IIRC) distribution if the residuals are normally distributed. They are both wald tests, they just differ in the distribution of the test statistic. If scipy underreports very small p-values, this would have a big effect on fields, where very small p-values are very important (like high-throughput genomic data). The two tests give very different results when the sample size is small and the slope is extreme, with Wald's test p-values being consistently lower than t-test's (even by orders of magnitude). Why is the p-value computed in such way? My go-to statistics textbook by Larry Wasserman, suggests that I compute the p-value of linear regression in a different way, by computing Wald Statistic, followed by a Wald test. It appears that scipy first computes the Wald Statistic on the estimator of the slope (gradient), followed by a (sampleSize - 2) degree of freedom t-test. # named tuple for scipy's linear regression

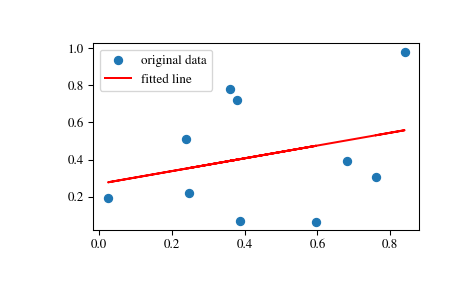
ResultT = LinregressResult(slope=B1, intercept=B0, rvalue=None, pvalue=pvalueT, stderr=stdB1) ResultWald = LinregressResult(slope=B1, intercept=B0, rvalue=None, pvalue=pvalueWald, stderr=stdB1) LinregressResult = collections.namedtuple("LinregressResult", ) # named tuple for our linear regression, with p-value coming from Wald Statistic PvalueT = 2*distributions.t.sf(abs(W), sampleSize - 2) StdB1 = errStd / (XStd * math.sqrt(sampleSize)) # Partial steps to compute Wald Statistic.ĮrrStd = math.sqrt((1/(sampleSize-2))*(sum())) # Partial steps to compute estimators of linear regression parameters.ī1 = sum(x * y for x, y in zip(XDiff, YDiff)) / sum(XDiffSquared)